Sir. William Bragg and Sir. Lawrence Bragg shared the Nobel Prize in 1915 for their work on analyzing crystal structures using X-Rays.
Father and Son Nobel Prize
This award is notable not only because it was the first and only time that a father-son duo has been awarded the Nobel but also because Lawrence was only 25 then. Indeed, he remained the youngest ever awardee until Malala Yousafzai’s (17) award for her work against the suppression of children’s education in 2014. Since the Bragg father-son award, a further 28 Nobel prizes have been awarded to X-ray crystallographers.
The Braggs first heard of the intriguing diffraction of X-rays by crystals in the summer of 1912. William Bragg received a letter describing a lecture by Max von Laue in which he demonstrated that X-rays were indeed waves rather than particles. He showed that when X-rays were passed through a crystal of Copper Sulphate, they produced wave-like diffraction patterns on photographic film, a discovery that won him the Nobel Prize in 1914. Henri Poincare, a French polymath, foresaw this observation's importance, though he did not live to see his prediction come true.
___
“if the value of discovery is to be measured by the fruitfulness of its consequences, the work of Laue and his collaborators should be considered as perhaps the most important of modern physics“
Henri Poincare (1912)
___
Beginnings for X-Ray Crystallography
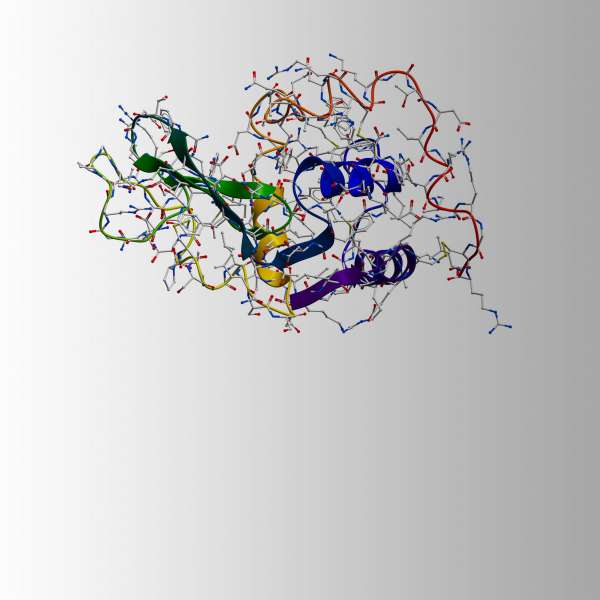
Lawrence Bragg, the duo's son, was convinced that von Laue’s initial explanation for the pattern of diffracted spots was flawed. He noticed that the diffracted spots became elliptical as the film was moved further away from the crystal. His explanation is now known as Bragg’s law, an equation describing the relationship between the pattern of spots, the wavelength of the light, and the deflected angle, with the planes of the atoms in the crystal. For us non-Nobel awardees, it was proven that you could calculate the orientation and organization of the individual atoms of a molecule using the pattern of spots on the X-ray film. Using this insight, the structures of simple crystals such as table salt (NaCl) and Potassium Chloride (KCl) were soon solved, leading the way for more complex molecules, including DNA in 1953 and Lysozyme in 1965.
___
“When I applied this test to Laue pictures, it was at once clear that all the spots were indeed in positions explained by reflection in the crystal planes. As a check, I set up an experiment in which a fine beam of X-rays fell upon a sheet of mica. Mica cleaves so readily that one would expect the sheets of atoms parallel to the cleavage to be well-marked. I found that for any angle of incidence of the rays, the mica always gave a reflected beam as if it were a mirror reflecting the radiation. I vividly remember taking my plate, still wet from the fixing dish, down to J. J. Thomson's room and showing it to him. It was very gratifying to see my professor's great excitement.“
___
The ability to deduce the 3-dimensional shape, that is, its structure based on the orientation of the atoms within a molecule, is made possible by the strict, symmetrical organization of crystals. So it follows that the first step required to understand the 3D shape of a molecule is successfully turning it into a crystal, a process called crystallization. Generally, the simpler the molecule, the simpler it is to crystallize. Whereas NaCl is composed of two atoms, biological molecules are much more complex - Lysozyme has nearly 2000. This makes the calculations and works required to solve the structure of proteins significantly harder.
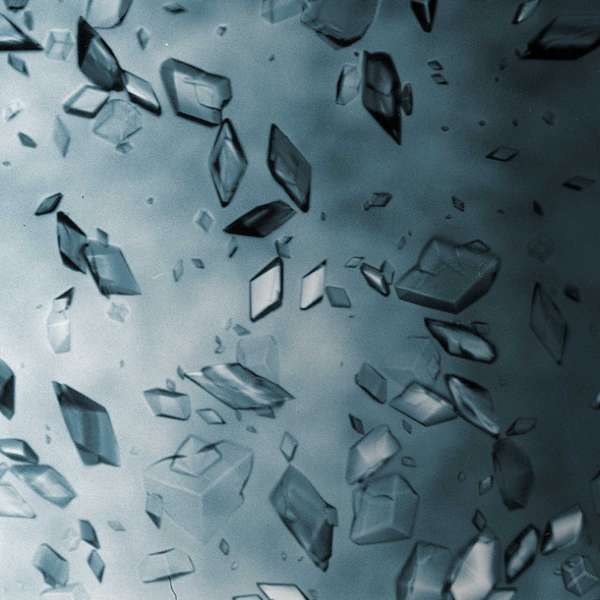
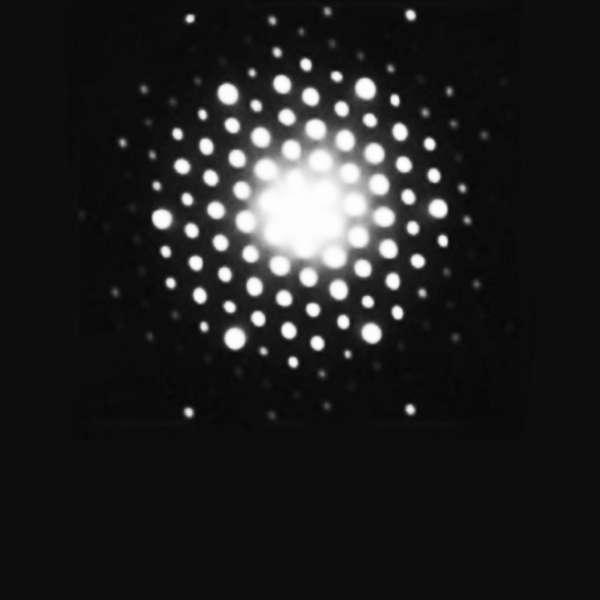
A simple salt such as Zincblende (Fig. 1) results in several diffracted X-ray spots on a photographic film. By rotating the crystal by a known amount and recording the intensity and position of the resulting spots, the position of electrons within the atoms of the molecule can be calculated. This overlayed stack of images produces what is referred to as an electron density map. There are of course, further complexities to making sure that alignment is maintained and that the positioning across multiple images is in-phase, but we will leave those confusions for elsewhere!
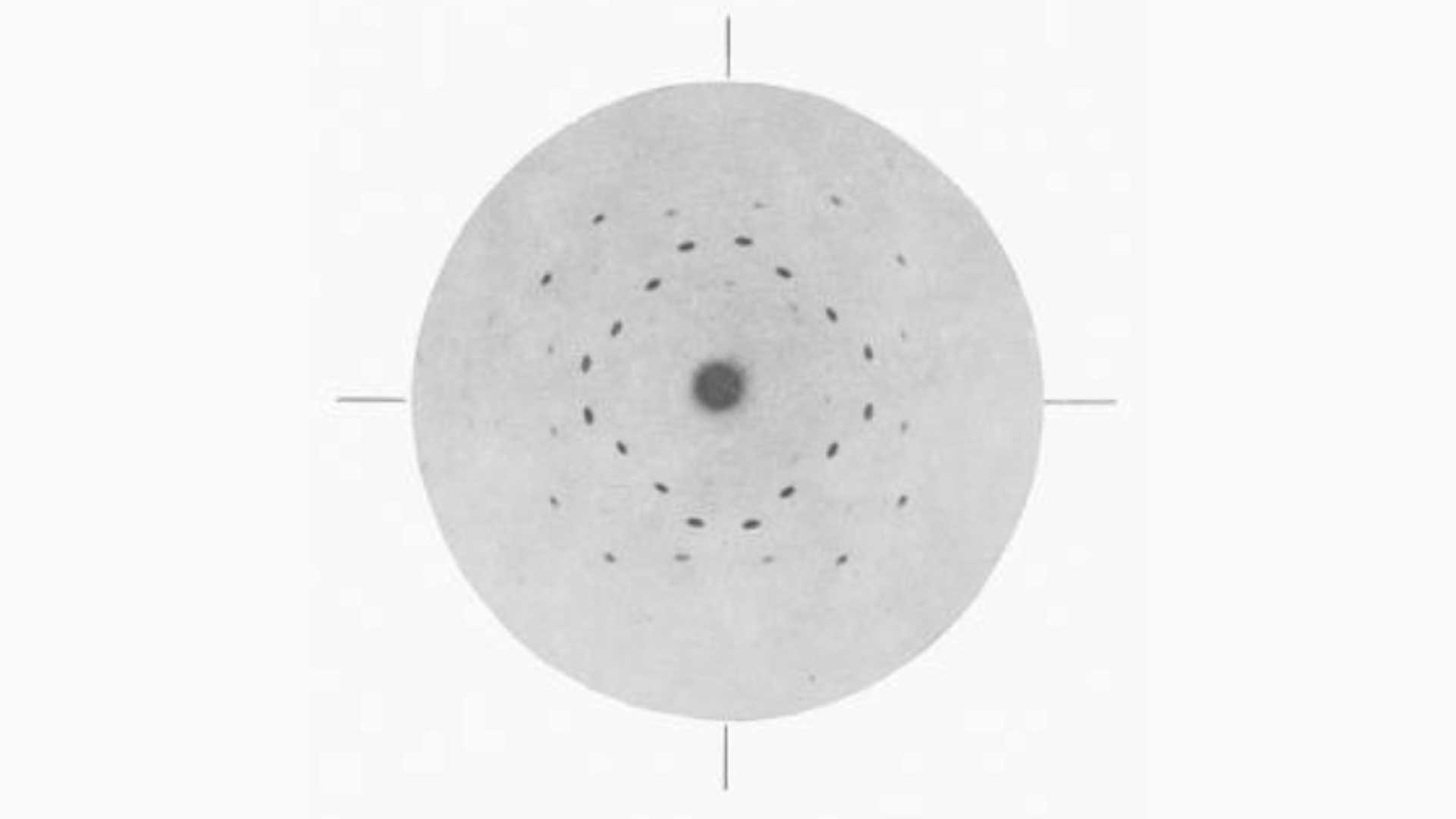

Fig. 1: Presentation of the solved structure of Lysozyme at the Royal Society in 1965. Courtesy of Louise Johnson, Oxford University.
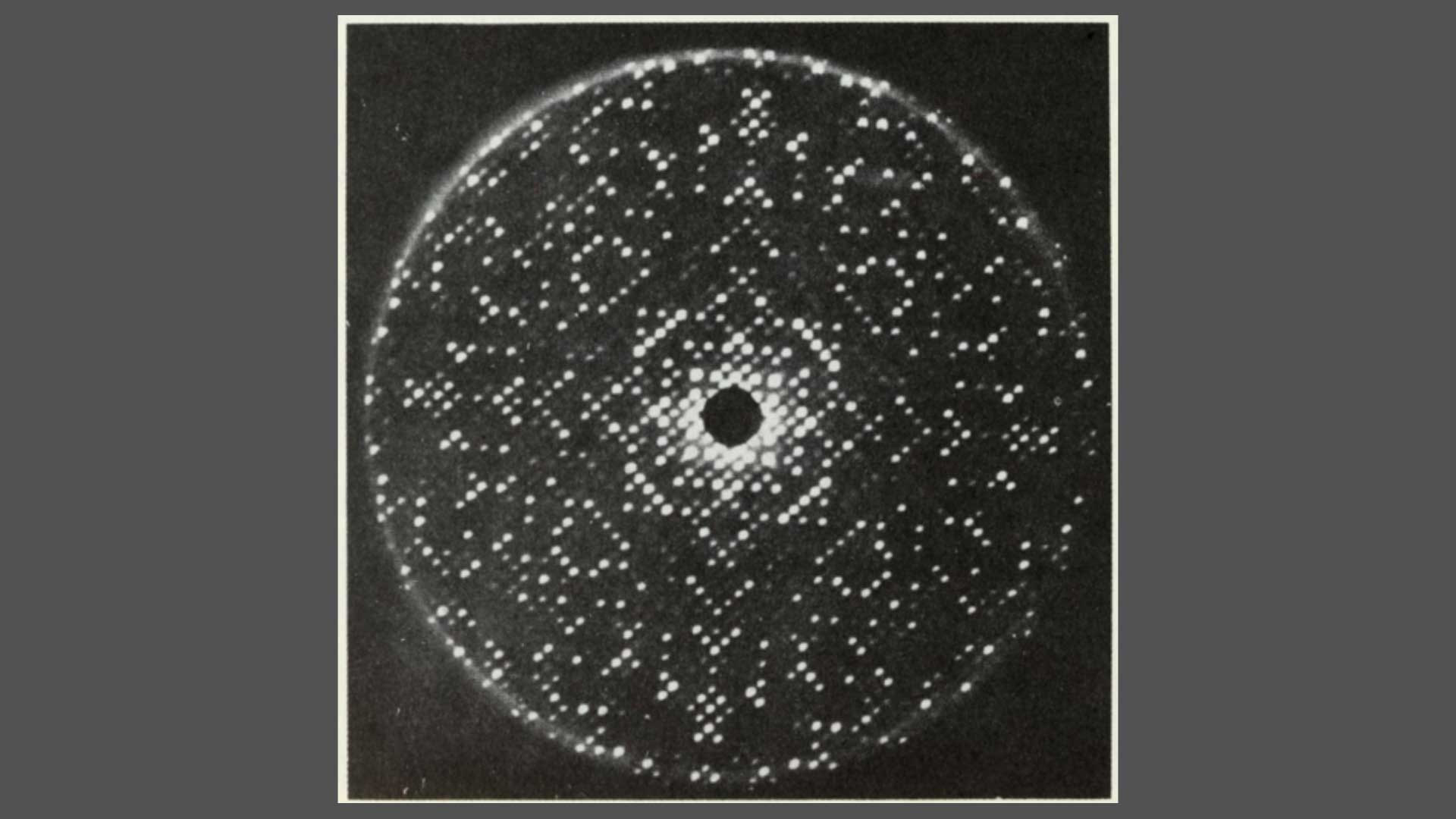

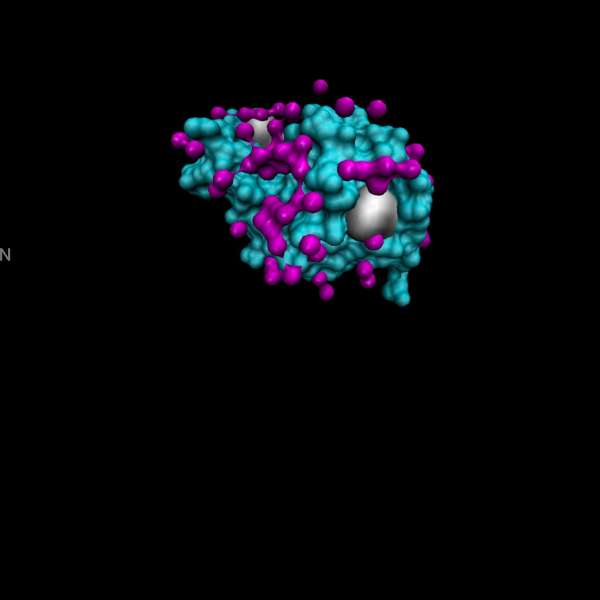
Fig. 2: The 2 Angstrom resolution electron density map. Sections Z = 25-34/60. The densities correspond to the number of amino-acid sidechains. From Louise Johnson, Science Progress 1966.
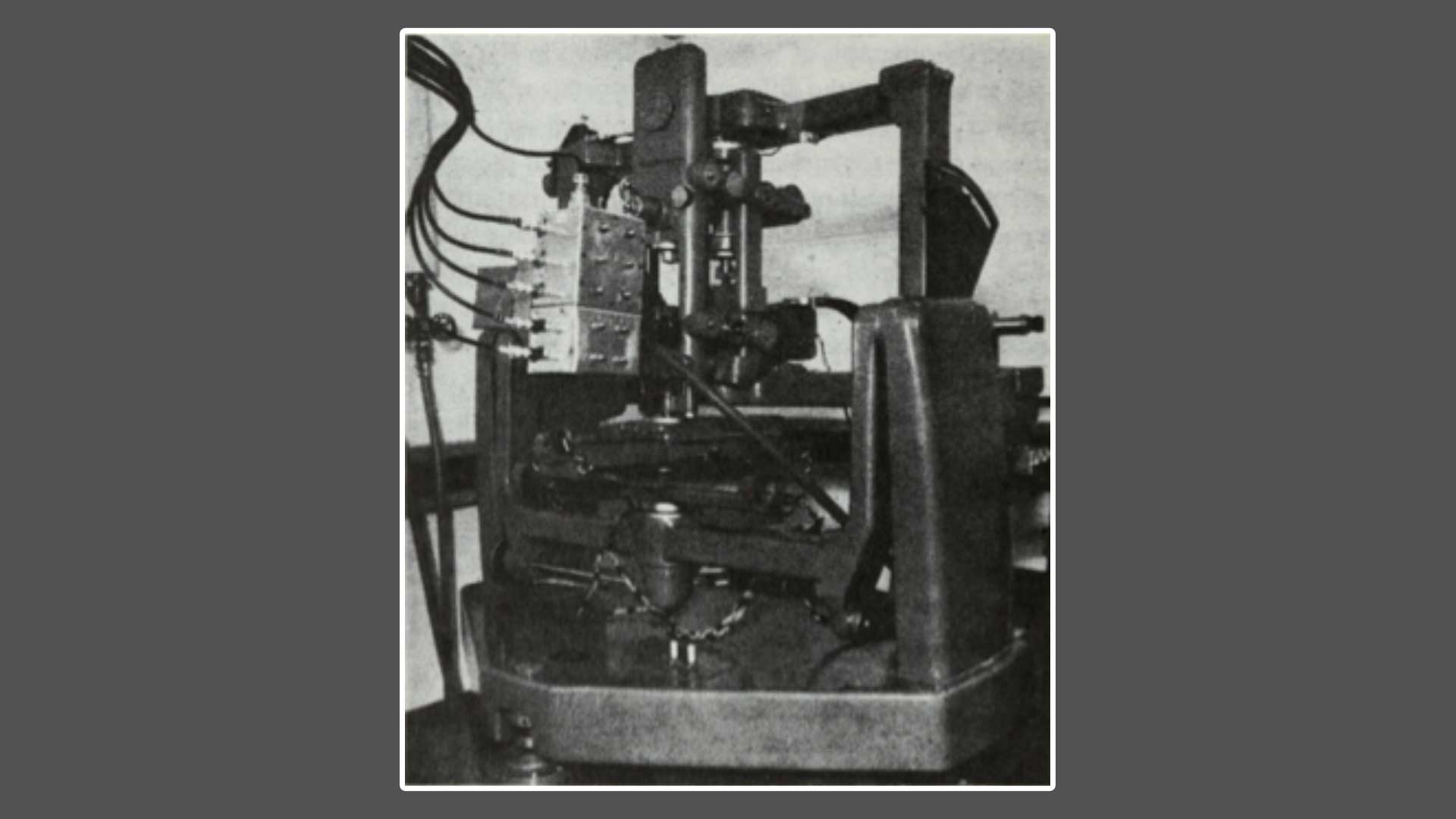

Fig. 3: Schematic drawing of the conformation of Lysozyme, drawn by Sir Lawrence Bragg From Louise Johnson, Science Progress 1966
Lysozyme: The third protein and first enzyme to be solved
___
“If the structure of a few molecules of a new type can be analyzed, a rich harvest is then reaped because the structures of many others will then be clear by analogy. I foresee the same happening in the protein field.“
Excerpt from Lawrence Bragg’s application for Medical Research Council Funding, 1947
___
The first protein structure solved was whale muscle myoglobin in 1957, earning John Kendrew and Max Perutz the 1962 Nobel Prize. That decade saw fewer than a dozen further protein structures solved due to the incredible complexity of and time required for this technique. In this context, work began solving the structure of Lysozyme in 1960, which had first been crystallized in 1937 by EP Abraham and R. Robinson. Luckily, two independent well-agreeing amino acid sequences had already been published by both the Jolles and Canfield groups, arming Lawrence Bragg’s team with a reference from which to work back.
X-ray generators, linear diffractometers, and a manually operated 3-circle diffractometer were used in the laboratory to solve Lysozyme’s structure. Crystals were painstakingly placed, rotated, images captured, calculations made, and the process repeated. Calculations were done either by hand or using the early Ferranti Mercury computer to which the group had access at night. Once electron density maps were in hand, two teams worked in shifts full-time for an entire month to align the amino acid sequence with the puzzle of the electron density map. An array of equally spaced grid points were drawn onto transparency sheets (similar to those used for overhead projectors, remember those?), and electron densities were plotted on these in series. These dots were connected by drawing contours through points of equal density, and the resulting 10,000 sheets were stacked to enable viewing of the 3D structure from above. This stack of transparency sheets could then be used to build the physical 3D model.
Work on the model was completed in time to serve as a centerpiece at William Bragg’s 75th birthday celebration. The structure was presented publicly for the first time at the Royal Institution on the 5th of November 1965. Luckily for us, photographs from this lecture exist and can be seen below. On the table stands the solved structure of Lysozyme, next to the simple NaCl crystal structure. Both are dwarfed by the 10m long unfolded amino-acid chain of Lysozyme, which extends beyond the photo toward the ceiling from which it hangs. The folded Lysozyme structure, built at the same scale as that of the unfolded 129 amino acid chain (2 cm per 2 Angstroms) illustrates the remarkable ability of proteins to take on compact functional forms once folded. In front of the desk is a Perspex box, within which stacked transparency sheets with contour-mapped electron densities are held. It is from these stacked sheets that the structure of Lysozyme was deduced. The model survives to this day in the Faraday Museum in the basement of the Royal Institution.
___
“The initiation of a project to study a selected protein structure requires very careful consideration; it is rather like a decision as to which new type of airplane to build. The cost of manpower, time, and money is considerable. Lysozyme, which Dr. Poljak had already studied when he joined the Davy Faraday team in 1960, proved to be a fortunate choice. It is the third protein structure to be successfully analyzed and the first enzyme. I have been forced to eat my words in two ways. I estimated not long ago that a protein analysis required fifty man-years of first-rate researchers' time and a quarter of a million pounds. The cost for lysozyme has been about one-half in each respect.“
Lawrence Bragg 1967: Excerpt from Proceedings of the Royal Society of London. Series B, Biological Sciences, Vol. 167, No. 1009, A Discussion on the Structure and Function of Lysozyme (Apr. 18, 1967)
___
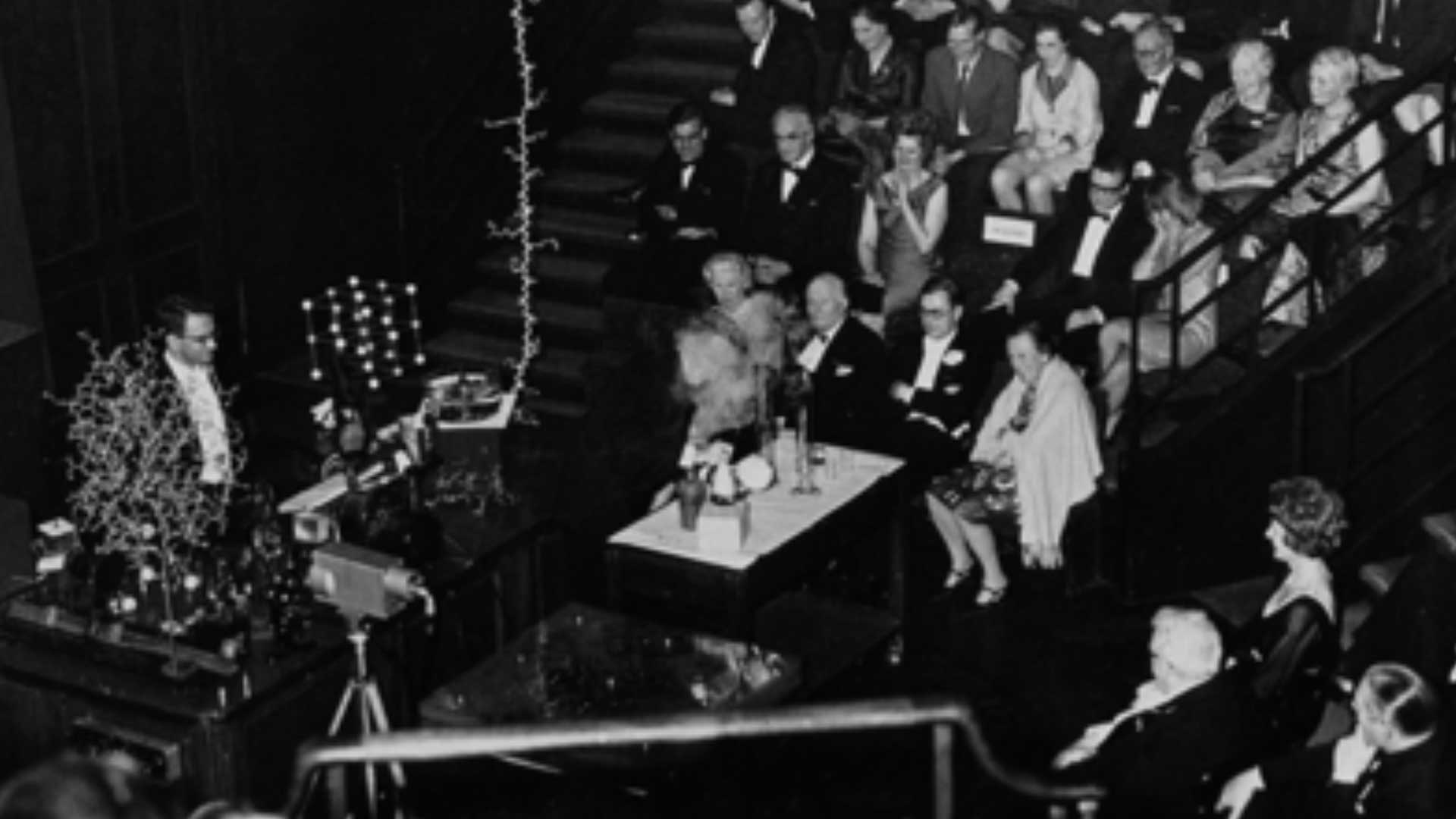

Fig. 4: Presentation of the solved structure of Lysozyme at the Royal Society in 1965. Courtesy of Louise Johnson, Oxford University.
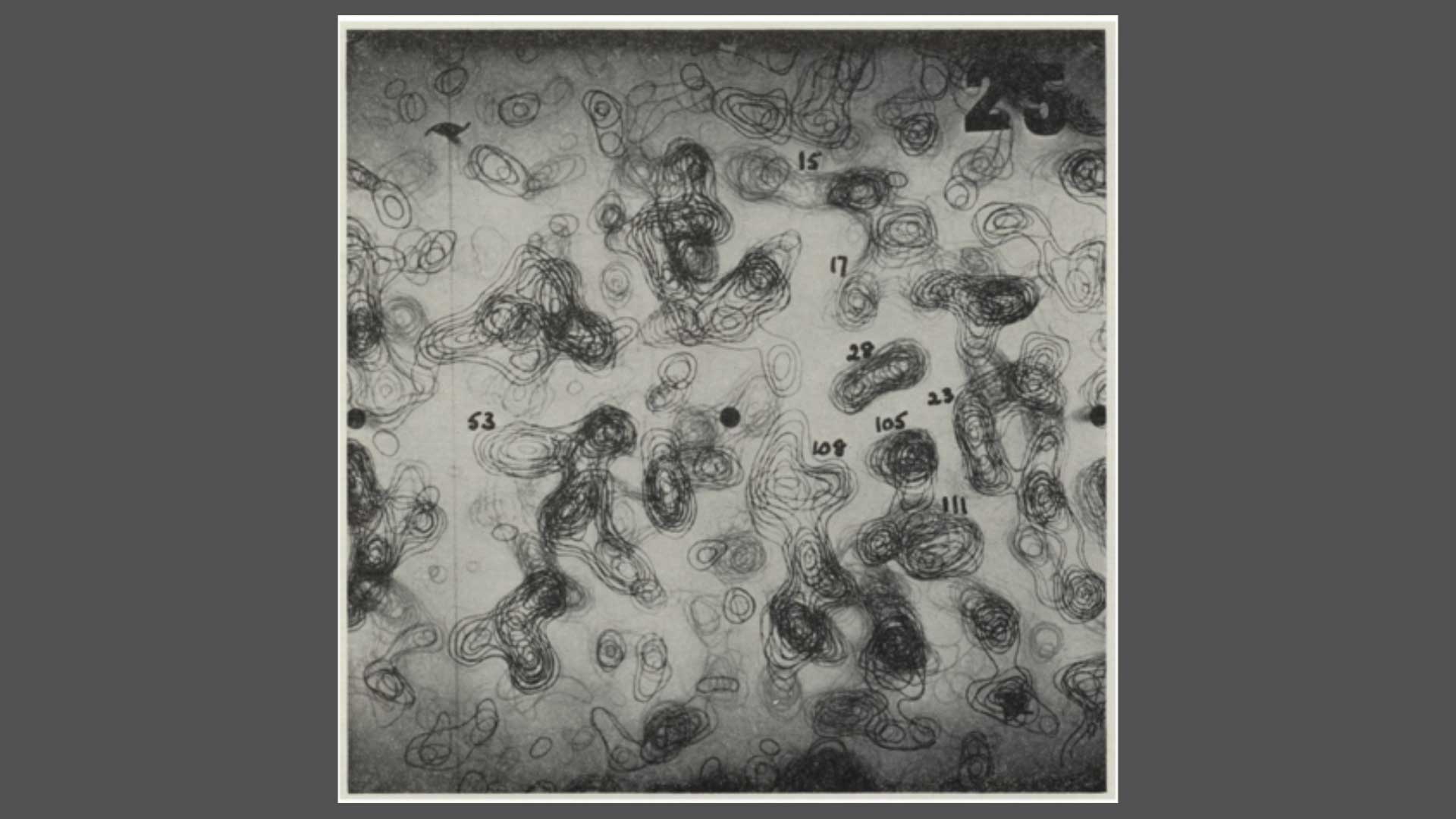

Fig. 5: The 2 Angstrom resolution electron density map. Sections Z = 25-34/60. The densities correspond to the number of the amino-acid sidechains. From Louise Johnson, Science Progress 1966.
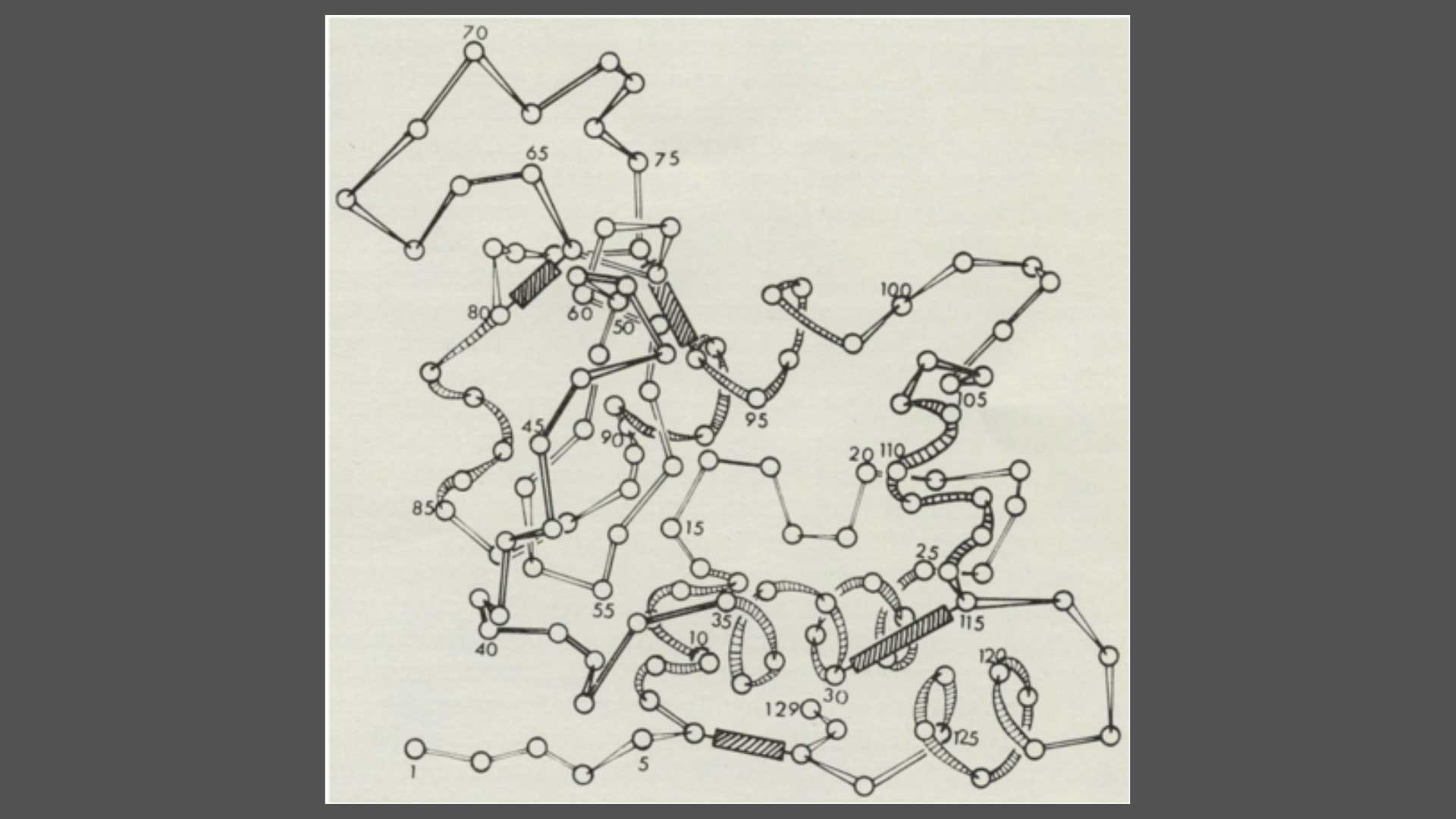
Fig. 6: Schematic drawing of the conformation of Lysozyme, drawn by Sir Lawrence Bragg From Louise Johnson, Science Progress 1966
100 years of Lysozyme
2022 marks 100 years since news of Lysozyme’s discovery reached the Royal Society. Let us take you on the journey of this 100-year story: